ARN Board: What is the positive evidence for evolution #6?
Rafe Gutman posted the following on ARN Board: What is the positive evidence for evolution #6?
(Also 'enjoy' the response by ID's Bulldog (vacuity of ID at work))
Read more!
(Also 'enjoy' the response by ID's Bulldog (vacuity of ID at work))
o how do we apply this to biology? if you were to compare the DNA sequence of a human gene, and a homologous mouse gene, they would probably be different by several nucleotides. for now let's restrict our analysis to the differences that do not result in a different amino acid, in other words, silent mutations. now imagine if you were to compare every human gene to its mouse counterpart, and score the number of silent mutations for each pair. what would you expect to see if the mutations were random? you would expect to see a distribution very much like a bell curve, which is in fact what we do see.
-
(note: this graph wasn't generated exactly how i described above, it's just to give you an idea. see here for the actual article)
you can apply this to any two organisms and get a bell curve out of it, with the average difference (i.e. peak of the curve) changing depending on the relatedness of the two organisms.
it's the success of these predictions from randomness that provide evidence for evolution, common ancestry, and RM+NS as the mechanism.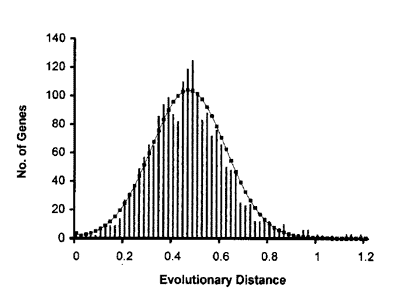
<< Home